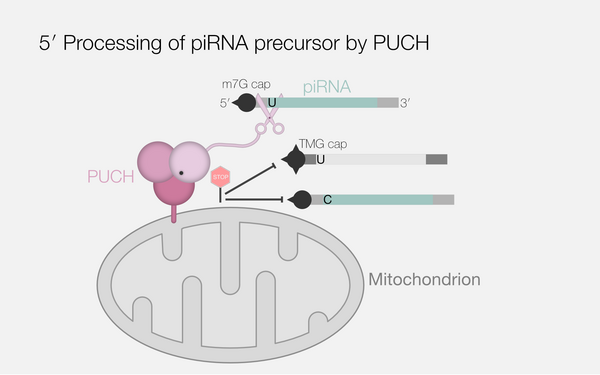
piRNA processing by a trimeric Schlafen-domain nuclease.
2023 NATURE;622(7982):402-409.
PMID: 37758951
Podvalnaya, Nadezda; Bronkhorst, Alfred W; Lichtenberger, Raffael; Hellmann, Svenja; Nischwitz, Emily; Falk, Torben; Karaulanov, Emil; Butter, Falk; Falk, Sebastian; Ketting, René F
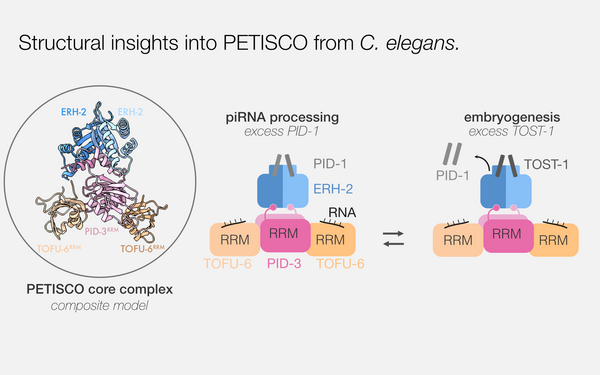
Structural basis of PETISCO complex assembly during piRNA biogenesis in C. elegans
2021 GENE DEV;35(17-18):1304-1323.
PMID: 34413138
Perez-Borrajero, Cecilia; Podvalnaya, Nadezda; Holleis, Kay; Lichtenberger, Raffael; Karaulanov, Emil; Simon, Bernd; Basquin, Jérôme; Hennig, Janosch; Ketting, René F; Falk, Sebastian
A ribonuclease III involved in virulence of Mucorales fungi has evolved to cut exclusively single-stranded RNA.
2021 NUCLEIC ACIDS RES
PMID: 33877360
Cánovas-Márquez, José Tomás; Falk, Sebastian; Nicolás, Francisco E; Padmanabhan, Subramanian; Zapata-Pérez, Rubén; Sánchez-Ferrer, Álvaro; Navarro, Eusebio; Garre, Victoriano
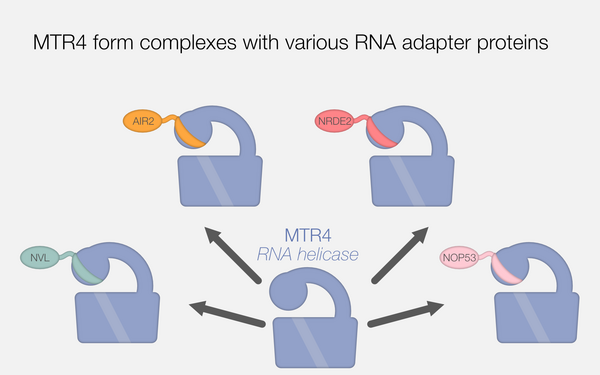
The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs
2019 NAT COMMUN
PMID: 31358741
Lingaraju, Mahesh; Johnsen, Dennis; Schlundt, Andreas; Langer, Lukas M.; Basquin, Jérôme; Sattler, Michael; Jensen, Torben Heick; Falk, Sebastian; Conti, Elena
Structure of the nuclear exosome captured on a maturing preribosome.
2018 SCIENCE;360(6385):219-222.
PMID: 29519915
Schuller, Jan Michael; Falk, Sebastian; Fromm, Lisa; Hurt, Ed; Conti, Elena
Mpp6 Incorporation in the Nuclear Exosome Contributes to RNA Channeling through the Mtr4 Helicase.
2017 Cell Rep;20(10):2279-2286.
PMID: 28877463
Falk, Sebastian; Bonneau, Fabien; Ebert, Judith; Kögel, Alexander; Conti, Elena
Structural insights into the interaction of the nuclear exosome helicase Mtr4 with the preribosomal protein Nop53.
2017 RNA;23(12):1780-1787.
PMID: 28883156
Falk, Sebastian; Tants, Jan-Niklas; Basquin, Jerôme; Thoms, Matthias; Hurt, Ed; Sattler, Michael; Conti, Elena
Structure of the RBM7-ZCCHC8 core of the NEXT complex reveals connections to splicing factors.
2016 NAT COMMUN;7:13573.
PMID: 27905398
Falk, Sebastian; Finogenova, Ksenia; Melko, Mireille; Benda, Christian; Lykke-Andersen, Søren; Jensen, Torben Heick; Conti, Elena
The molecular architecture of the TRAMP complex reveals the organization and interplay of its two catalytic activities.
2014 MOL CELL;55(6):856-867.
PMID: 25175027
Falk, Sebastian; Weir, John R; Hentschel, Jendrik; Reichelt, Peter; Bonneau, Fabien; Conti, Elena