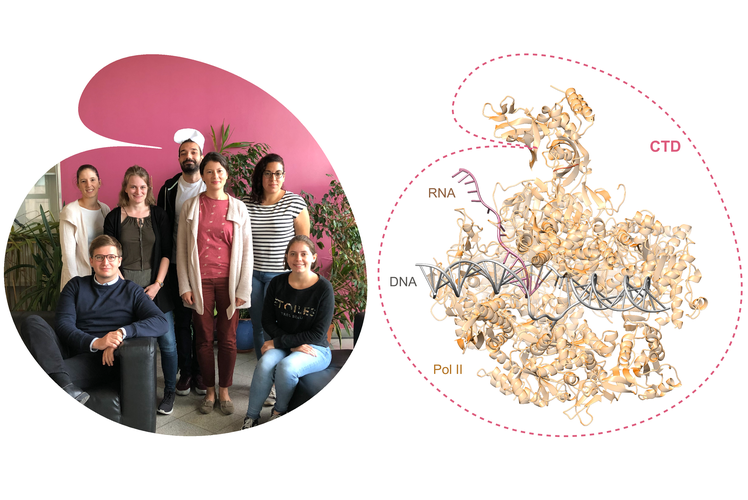
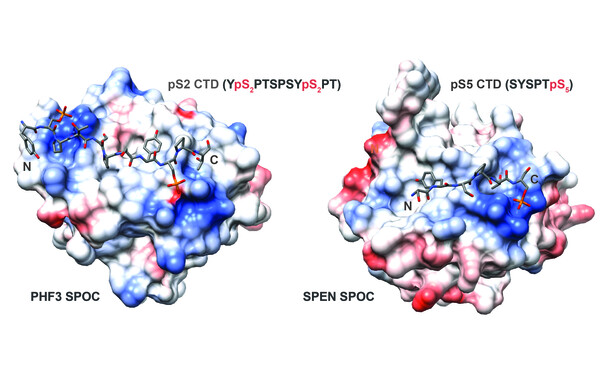
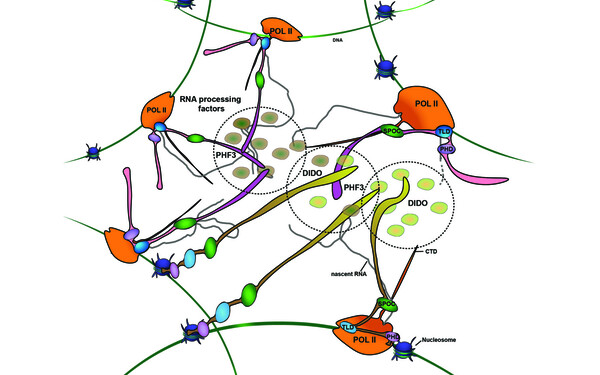
PHF3 regulates neuronal gene expression through the Pol II CTD reader domain SPOC.
2021 NAT COMMUN;12(1):6078.
PMID: 34667177
Appel, Lisa-Marie; Franke, Vedran; Bruno, Melania; Grishkovskaya, Irina; Kasiliauskaite, Aiste; Kaufmann, Tanja; Schoeberl, Ursula E; Puchinger, Martin G; Kostrhon, Sebastian; Ebenwaldner, Carmen; Sebesta, Marek; Beltzung, Etienne; Mechtler, Karl; Lin, Gen; Vlasova, Anna; Leeb, Martin; Pavri, Rushad; Stark, Alexander; Akalin, Altuna; Stefl, Richard; Bernecky, Carrie; Djinovic-Carugo, Kristina; Slade, Dea
The SPOC domain is a phosphoserine binding module that bridges transcription machinery with co- and post-transcriptional regulators.
2023 NAT COMMUN;14(1):166.
PMID: 36631525
Appel, Lisa-Marie; Franke, Vedran; Benedum, Johannes; Grishkovskaya, Irina; Strobl, Xué; Polyansky, Anton; Ammann, Gregor; Platzer, Sebastian; Neudolt, Andrea; Wunder, Anna; Walch, Lena; Kaiser, Stefanie; Zagrovic, Bojan; Djinovic-Carugo, Kristina; Akalin, Altuna; Slade, Dea
The SPOC proteins DIDO3 and PHF3 co-regulate gene expression and neuronal differentiation.
2023 NAT COMMUN;14(1):7912.
PMID: 38036524
Benedum, Johannes; Franke, Vedran; Appel, Lisa-Marie; Walch, Lena; Bruno, Melania; Schneeweiss, Rebecca; Gruber, Juliane; Oberndorfer, Helena; Frank, Emma; Strobl, Xué; Polyansky, Anton; Zagrovic, Bojan; Akalin, Altuna; Slade, Dea
SPOC domain proteins in health and disease.
2023 GENE DEV
PMID: 36927757
Appel, Lisa-Marie; Benedum, Johannes; Engl, Magdalena; Platzer, Sebastian; Schleiffer, Alexander; Strobl, Xué; Slade, Dea
A novel non-canonical PIP-box mediates PARG interaction with PCNA
2017 NUCLEIC ACIDS RES
PMID: 28934471
Tanja Kaufmann, Irina Grishkovskaya, Anton A. Polyansky, Sebastian Kostrhon, Eva Kukolj, Karin M. Olek, Sebastien Herbert, Etienne Beltzung, Karl Mechtler, Thomas Peterbauer, Josef Gotzmann, Lijuan Zhang, Markus Hartl, Bojan Zagrovic, Kareem Elsayad, Kristina Djinovic-Carugo, Dea Slade
PARP inhibition causes premature loss of cohesion in cancer cells.
2017 Oncotarget;8(61):103931-10395.
PMID: 29262611
Kukolj, Eva; Kaufmann, Tanja; Dick, Amalie E; Zeillinger, Robert; Gerlich, Daniel W; Slade, Dea
PARP and PARG inhibitors in cancer treatment.
2020 GENE DEV;5-6(34):360-394.
PMID: 32029455
Slade, Dea
SIRT2 regulates nuclear envelope reassembly via ANKLE2 deacetylation.
2016 J CELL SCI(129): 4607-4621.
PMID: 27875273
Kaufmann, Tanja; Kukolj, Eva; Brachner, Andreas; Beltzung, Etienne; Bruno, Melania; Kostrhon, Sebastian; Opravil, Susanne; Hudecz, Otto; Mechtler, Karl; Warren, Graham; Slade, Dea
Direct measurement of protein-protein interactions by FLIM-FRET at UV laser-induced DNA damage sites in living cells.
2020 NUCLEIC ACIDS RES;21(48)
PMID: 33053171
Kaufmann, T; Herbert, S; Hackl, B; Besold, J M; Schramek, C; Gotzmann, J; Elsayad, K; Slade, D