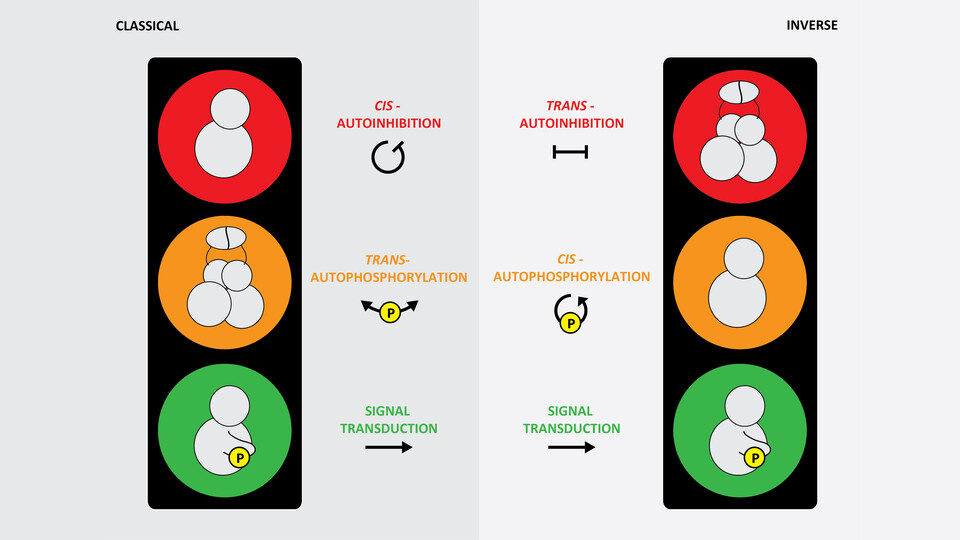

Of the approximately 10000 proteins in a typical mammalian cell, about a third are phosphorylated at any given time. This reversible chemical modification controls the activity and localization of proteins and is therefore a key regulatory step in many cellular processes, from cell signaling to metabolism. The Leonard lab focuses on the molecular mechanisms of signal transduction. As part of their work, the scientists study protein kinases, enzymes that catalyze phosphorylation and, thereby, transduce and propagate signals within cells. In 2019, the Leonard lab discovered a ubiquitin-like domain (ULD) in PKD that mediates its dimerization. Since PKD was known to require phosphorylation in order to be activated, the scientists assumed that PKD must phosphorylate its own activation loop in trans. This dimerization-mediated trans model of kinase autoactivation has been observed in many auto-phosphorylating protein kinases, and thus seemed a logical conclusion.
But when PhD student Ronja Reinhardt started working on the project, she soon faced major challenges: "We had a clear hypothesis, but for a long time our assays didn’t work out the way we thought they would and the data just didn’t add up." The scientists soon figured out why. Counterintuitively, autophosphorylation of PKD was inhibited by dimerization. "Through a lot of critical thinking and careful experimentation, we came to understand that PKD is actually activated in completely the inverse mechanism to what we expected." In the scientists’ revised model, PKD is by default dimeric and locked in a compact, inactive conformation. When it binds to the membrane signaling lipid diacylglycerol, it undergoes conformational changes that force the kinase domains to dissociate from one another, thereby freeing them to phosphorylate themselves in cis. This means that each kinase domain phosphorylates its own activation loop.
The mechanistic details of the process that releases the protein from its locked conformation remain to be elucidated. The study also raises questions about the activation mechanisms of other protein kinases. "For me the most exciting aspect of the work is that it reveals a completely unexpected but extremely logical mechanism that nature has found to solve the same problem with a different solution", says group leader Thomas Leonard. The work also teaches a lesson about the scientific process itself, which the researchers are keen to point out: "Our work illustrates the importance of rigorously testing scientific hypotheses. A negative result is just as valuable as a positive one, and one should always rethink the hypothesis based on the experimental data, and not the other way around," Thomas Leonard concludes.
Ronja Reinhardt, Kai Hirzel, Gisela Link, Stephan A Eisler, Tanja Hägele, Matthew A H Parson, John E Burke, Angelika Hausser, Thomas A Leonard: PKD autoinhibition in trans regulates activation loop autophosphorylation in cis. PNAS 2023