Vaccinia virus, the virus that served as the vaccine to eradicate smallpox, is studied extensively as a model for virus-host interactions, as it employs a whole arsenal of mechanisms to interfere with the hosts immune system. Of its almost 200 genes only half encode proteins required for viral replication. Many of the other genes encode proteins such as A46 that directly interfere with the immune system of the infected organism. “We knew that A46 interacted with several players of the immune system, but the mechanisms of interaction of A46 with its targets has remained unclear”, explains group leader Tim Skern.
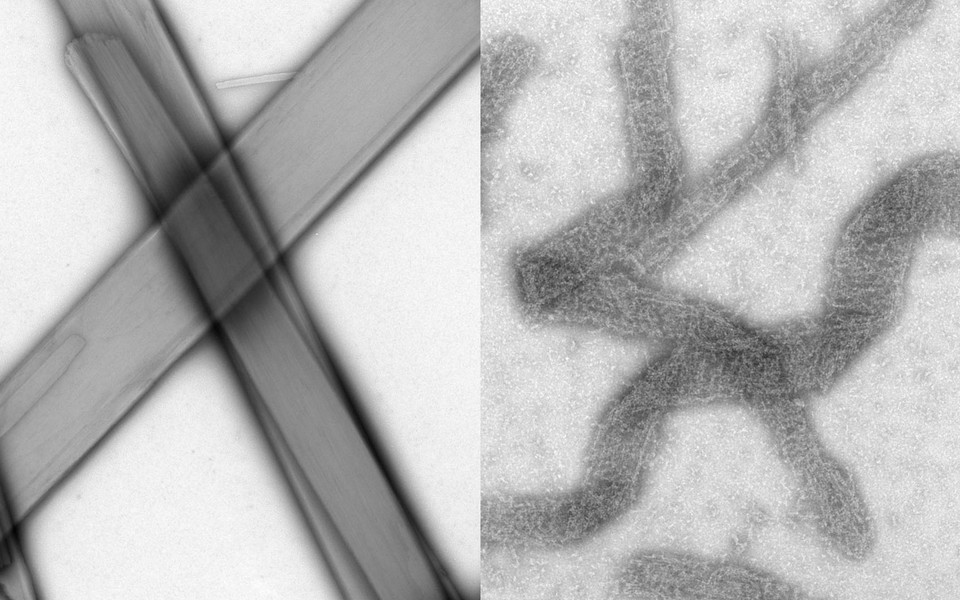
Applying electron microscopy, structural biology, and protein crosslinking techniques, PhD student Daniel Azar and Master’s student Meryl Haas now show that A46 destroys assemblies formed by two proteins called MyD88 and MAL. These intracellular adaptor proteins dock to receptors in the cell membrane and relay immune signals to the nucleus where antiviral gene expression is activated. Both proteins are known to form cable-like filaments that transduce these signals. Filament formation of MyD88 and MAL is driven by the interaction of their TIR domains.
The researchers identified the patches in A46 that target the TIR domains of MyD88 and MAL. Based on their experimental findings, the scientists propose that A46 intercalates into the filaments formed by the host proteins and thereby disrupts the signaling chain. “This is an unusual mechanism for viral proteins as it retains the basic structure and function of the host proteins”, says Tim Skern. “More commonly, host proteins are cleaved by the pathogen, rendering them useless”. Moreover, A46 exerts its destructive force in a concentration dependent manner. At lower ratios of A46 to MAL or MyD88, the protein affects filaments by chopping and unwinding them. At higher concentrations of A46, filament assemblies are completely destroyed.
Viruses have a genome ranging from only a few to hundreds of genes, yet manage to propagate and even take over organisms with a much larger genome. “How they manage to do so is a fundamental question in virology that my lab has been tackling for years. Our study now provides insight into a previously unknown mechanism of signaling inhibition.” concludes Tim Skern.
"Science Flash" presentation by first author Daniel Azar explaining the findings.
Original Publication:
Daniel F. Azar, Meryl Haas, Sofiya Fedosyuk, Md. Habibur Rahaman, Andrew Hedger, Bostjan Kobe and Tim Skern: Vaccinia virus immunomodulator A46: Destructive interactions with MAL and MyD88 shown by negative-stain EM
https://www.cell.com/structure/fulltext/S0969-2126(20)30332-4