
Stem cells have the unique property of being able to give rise to all the different cell types of an organism. This process of differentiation is governed by dramatic changes in gene expression patterns. The Leeb lab studies the molecular basis of these cell fate choices in mouse embryonic stem cells. Previous work from the lab resulted in a comprehensive catalogue of genes involved in the exit from naïve pluripotency. “One of the most surprising findings in these screens were components of the NMD pathway”, says group leader Martin Leeb. “This pathway is primarily known to eliminate aberrant mRNA transcripts that are potentially harmful to the cell, but a direct involvement in cell fate determination was unexpected”.
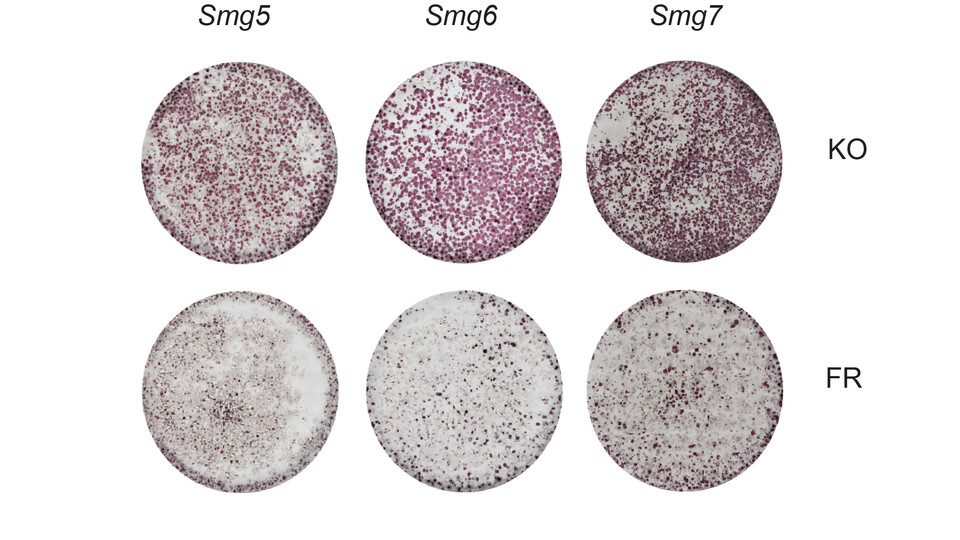
The scientists, including first authors Michelle Huth and Laura Santini, set out to characterize precisely how NMD affects transcription networks during differentiation. Using SLAM-seq technology developed by the lab of Stefan Ameres, they identified direct mRNA targets of the key NMD effector proteins SMG5, SMG6 and SMG7. Knockouts of these proteins caused increased translation rates and elevated levels of naïve pluripotency specific transcripts. “Our findings establish a hitherto unexpected role for NMD in differentiation”, says Martin Leeb. “However, it makes sense that, in order to establish a new cell state, the transcriptome of the previous one first needs to be erased. NMD may, therefore, help to create a ‘blank slate’ for the new cell identity.”
Consistent with the increased translation rates in NMD deficient ES cells, the researchers identified the translation initiation factor eIF4A2 as a novel NMD target gene. NMD disruption leads to the production of a truncated isoform of eIF4A2, which results in elevated activity of mTORC1 and increased translation rates. Translation itself is a fundamental requirement for NMD, but why the translation initiation factor eIF4A2 is a target of programmed decay by NMD remains enigmatic. The scientists hypothesize that there could be a previously unrecognized feedback mechanism at work that balances NMD with translational activity. “It is fascinating that, in the course of this work, we could not only learn something new about cell differentiation, but also about the process of nonsense mediated decay”, comments Michelle Huth.
Michelle Huth, Laura Santini, Elena Galimberti, Julia Ramesmayer, Fabian Titz-Teixeira, Robert Sehlke, Michael Oberhuemer, Sarah Stummer, Veronika Herzog, Marius Garmhausen, Merrit Romeike, Anastasia Chugunova, Friederike Leesch, Laurenz Holcik, Klara Weipoltshammer, Andreas Lackner, Christian Schoefer, Arndt von Haeseler, Christa Buecker, Andrea Pauli, Stefan L. Ameres, Austin Smith, Andreas Beyer, and Martin Leeb: NMD is required for timely cell fate transitions by fine-tuning gene expression and regulating translation. Genes & Development 2022.
doi: 10.1101/gad.347690.120