
Fungi, plants, and animals have more in common than first meets the eye. Unlike archaea and bacteria, these organisms are eukaryotes, meaning that they store their genome in a dedicated subcellular compartment called the nucleus. The compartmentalization of the genome both safeguards the DNA and permits more intricate regulatory control over gene expression, but it causes a logistical problem. In order to regulate gene expression as well as synthesize proteins, select molecules must be able to cross the double-layered nuclear envelope in both directions. Nuclear pore complexes are the gatekeepers to the genome and the cytoplasm.
On the molecular scale, these proteinaceous channels are giant. They consist of multiple copies of over 30 different proteins that form 6-8 subcomplexes within each pore. Hundreds of individual proteins are required to build just one NPC, of which there are thousands embedded in the nuclear envelope of a single cell. “Due to the complexity of nuclear pore complexes and the fleeting nature of the assembly events, it has been technically challenging to study the assembly of these structures”, explains first author Shotaro Otsuka, who pursued the project during his postdoctoral research in the Ellenberg Lab before becoming a group leader at the Max Perutz Labs. Consequently, scientists have struggled to understand how these complex molecular machines are built.
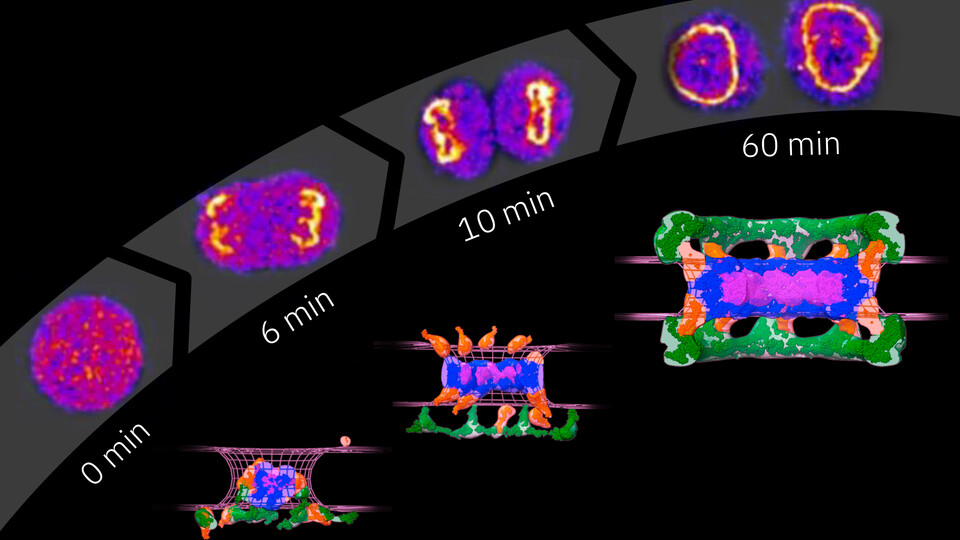
To overcome the challenges associated with studying the NPC, the scientists utilized a quantitative fluorescence imaging approach previously developed in the Ellenberg lab. The technique allows them to count the absolute number of proteins involved in a specific process in the cell over time. First, they fluorescently tagged nucleoporins, the basic building blocks of NPCs. Using microscopy, they then quantified how their copy number changes during the assembly of NPCs. To integrate their data with structural models of the assembling NPC that they had previously obtained, the team collaborated with the lab of Andrej Sali (University of California San Franscisco). Utilizing computational modeling tools and their quantitative information on protein copy numbers over time, the scientists were able to propose a detailed model of post-mitotic NPC assembly.
In higher eukaryotes NPCs assemble around the reforming nuclear envelope during mitotic cell division, as well as during the nuclear envelope expansion phase of the cell cycle. The scientists data also revealed key mechanistic differences between the two assembly pathways. During post-mitotic NPC assembly, components of the central ring build the inner core of the pore prior to addition of either cytoplasmic or nuclear filament proteins. However, in interphase assembly during nuclear envelope expansion, the central ring is added only after the cytoplasmic and nuclear filaments are assembled. A key remaining challenge will be to create a similarly detailed model for NPC assembly during interphase. “Beyond the NPC, our approach of combining quantitative fluorescence live-cell imaging with existing structural data could also prove a useful strategy in helping to understand the assembly mechanisms of other molecular machines in the cell”, concludes Shotaro Otsuka.
Publication:
Shotaro Otsuka, Jeremy O. B. Tempkin, Wanlu Zhang1 Antonio Z. Politi, Arina Rybina, M. Julius Hossain, Moritz Kueblbeck, Andrea Callegari, Birgit Koch, Natalia Rosalia Morero, Andrej Sali and Jan Ellenberg: A quantitative map of nuclear pore assembly reveals two distinct mechanisms. Nature 2023.
https://doi.org/10.1038/s41586-022-05528-w
https://www.nature.com/articles/s41586-022-05528-w