
Nuclear pores in the nuclear membrane do not only control the transport of molecules into and out of the nucleus but also play an important role in gene expression. Researchers at the Max F. Perutz Laboratories (MFPL) of the University of Vienna and the Medical University of Vienna have deciphered a mechanism by which nuclear pores use "interpreters" to directly influence gene expression. The findings recently published in Cell were a collaborative effort with scientists at the Research Institute of Molecular Pathology (IMP) and at the Pennsylvania State University (USA).
Scientists have been racking their brains over nuclear pores. Their role in the transport of molecules between nucleus and cytoplasm has been extensively described. Such molecules also include building blocks that help decipher the genetic information stored within the nucleus and guide those that have to leave the nucleus with a particular message out of it. This information is needed to produce proper proteins in the cytoplasm.
However, nuclear pores take an even more active role in gene expression, by directly modulating transcription of genetic information. „Genes are dynamic and change their position depending on if they are active or not. Intriguingly, some genes are able to attach to nuclear pores – the site where copies of genes are exported into the cytoplasm. This leads to changes in gene expression, in other words the frequency with which the transcriptional machinery reads genetic information. But how do nuclear pores communicate with genetic information? We had already assumed that specific adapters would not only attach genes to nuclear pores but that they would also pass on information to the transcriptional machinery,“ says Alwin Köhler, who is a group leader at the Max F. Perutz Laboratories.
In collaboration with researchers at the neighboring institute IMP in Vienna and colleagues at the Pennsylvania State University (USA), Alwin Köhlers group has succeeded in deciphering a mechanism that explains communication between nuclear pores and genes. „During our research, we discovered a ‚translation program’ that prepares the genetic information for RNA polymerase II,“ explains Alwin Köhler. RNA polymerase II is the key player during transcription and generates copies (messenger RNA, mRNA) that are necessary for protein synthesis.
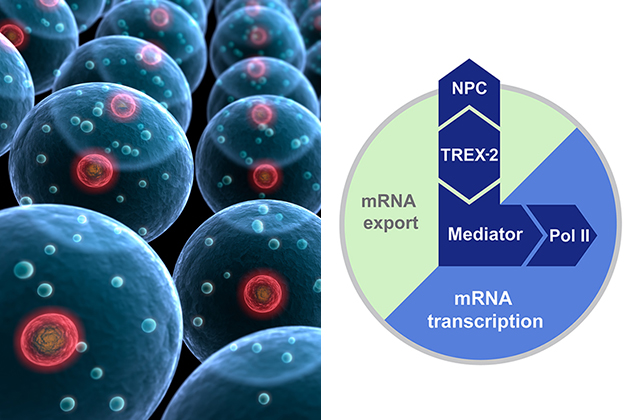
In detail, the researchers identified the so-called TREX-2 complex as interpreter number one. TREX-2 receives information directly from the nuclear pore and translates this message for a complex called Mediator. Subsequently, Mediator makes the information accessible to RNA polymerase II. The resulting mRNA is sent back to TREX-2 and the nuclear pore and then exported into the cytoplasm.
Following the publication in Developmental Cell in May, this is already the second study of Alwin Köhler’s group published in one of the most important scientific journals in the field of basic research this year. „Deciphering transcriptional mechanisms is crucial for understanding the blueprint for a cell. This lays the basis for eventually treating numerous diseases such as cancer, since diseases are often caused by 'communication errors' during gene expression. Ideally, basic and clinical research go hand in hand. At the Vienna Biocenter, the Medical University of Vienna has the great opportunity to work closely with excellent researchers of the University of Vienna and the neighboring institutes. The best example is our successful collaboration with Tim Clausen at the IMP,“ says Alwin Köhler. But the real work is only just beginning: „We have answered one question, however, ten new questions have arisen. That is precisely what makes this project so fascinating,“ adds Maren Schneider, one of the first authors of the study.
Publication in Cell:
Maren Schneider, Doris Hellerschmied, Tobias Schubert, Stefan Amlacher, Vinesh Vinayachandran, Rohit Reja, B. Franklin Pugh, Tim Clausen and Alwin Köhler: The nuclear pore-associated TREX-2 complex employs Mediator to regulate gene expression.