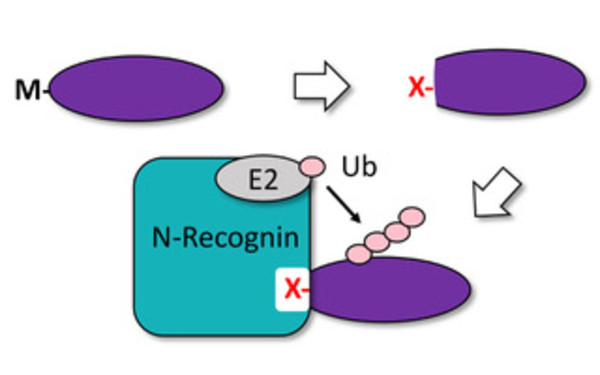
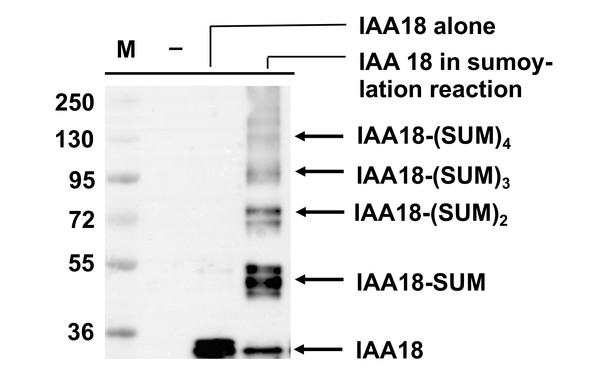
The UBR Domain of Plant Ubr1 Homolog PRT6 Accommodates Basic and Hydrophobic Amino Termini for Substrate Recognition.
2025 Journal of molecular biology;437(4):168939.
PMID: 39799992
Rudi Olga, Hodakova Zuzana, Farias Saad Carolina, Winter Nikola, Grishkovskaya Irina, Böhm Jessica, Jarck Greta, Schleiffer Alexander, Haselbach David, Bachmair Andreas
BIG enhances Arg/N-degron pathway-mediated protein degradation to regulate Arabidopsis hypoxia responses and suberin deposition.
2024 The Plant cell;36(9):3177, 3200, 3177-3200.
PMID: 38608155
Zhang Hongtao, Rundle Chelsea, Winter Nikola, Miricescu Alexandra, Mooney Brian C, Bachmair Andreas, Graciet Emmanuelle, Theodoulou Frederica L
A Yeast-Based Functional Assay to Study Plant N-Degron - N-Recognin Interactions.
2021 Frontiers in plant science;12:806129.
PMID: 35069663
Kozlic Aida, Winter Nikola, Telser Theresia, Reimann Jakob, Rose Katrin, Nehlin Lilian, Berckhan Sophie, Sharma Gunjan, Dambire Charlene, Boeckx Tinne, Holdsworth Michael J, Bachmair Andreas
Cellular Control of Protein Turnover via the Modification of the Amino Terminus.
2021 International journal of molecular sciences;22(7)
PMID: 33805528
Winter Nikola, Novatchkova Maria, Bachmair Andreas
The Scope, Functions, and Dynamics of Posttranslational Protein Modifications.
2019 Annual review of plant biology;70:119, 151, 119-151.
PMID: 30786234
Millar A Harvey, Heazlewood Joshua L, Giglione Carmela, Holdsworth Michael J, Bachmair Andreas, Schulze Waltraud X
Distinct branches of the N-end rule pathway modulate the plant immune response.
2019 The New phytologist;221(2):988, 1000, 988-1000.
PMID: 30117535
Vicente Jorge, Mendiondo Guillermina M, Pauwels Jarne, Pastor Victoria, Izquierdo Yovanny, Naumann Christin, Movahedi Mahsa, Rooney Daniel, Gibbs Daniel J, Smart Katherine, Bachmair Andreas, Gray Julie E, Dissmeyer Nico, Castresana Carmen, Ray Rumiana V, Gevaert Kris, Holdsworth Michael J
Sumoylation and phosphorylation: hidden and overt links.
2018 Journal of experimental botany;69(19):4583, 4590, 4583-4590.
PMID: 29846689
Tomanov Konstantin, Nukarinen Ella, Vicente Jorge, Mendiondo Guillermina M, Winter Nikola, Nehlin Lilian, Weckwerth Wolfram, Holdsworth Michael J, Teige Markus, Bachmair Andreas