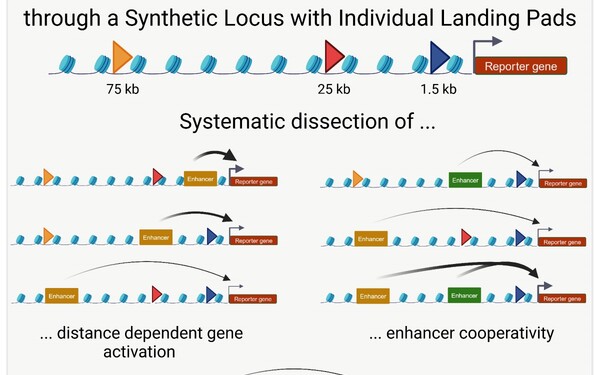
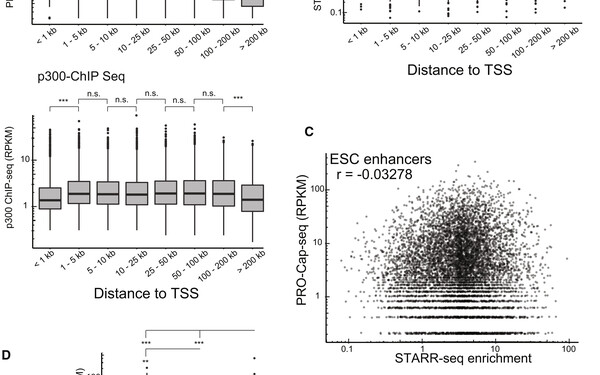
Enhancer cooperativity can compensate for loss of activity over large genomic distances.
2025 Molecular cell;85(2):362, 375.e9, 362-375.e9.
PMID: 39626663
Thomas Henry F, Feng Songjie, Haslhofer Felix, Huber Marie, García Gallardo María, Loubiere Vincent, Vanina Daria, Pitasi Mattia, Stark Alexander, Buecker Christa
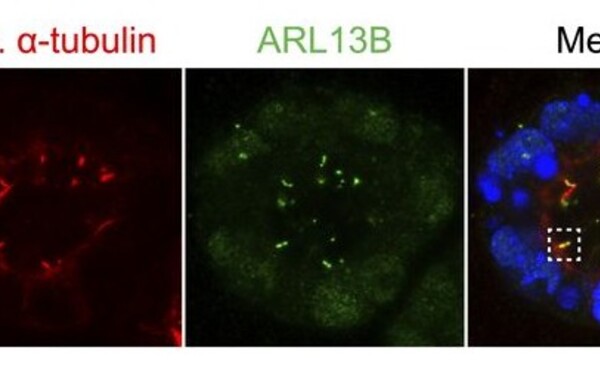
In vitro approaches to study centriole and cilium function in early mouse embryogenesis.
2026 Life science alliance;9(1)
PMID: 41161866
Voelkl Isabella, Civetta Tamara, Egg Mirijam, Huber Marie, Feng Songjie, Dammermann Alexander, Buecker Christa
The exit from naive pluripotency: a platform for the study of enhancer mechanistics.
2025 Biochemical Society transactions
PMID: 40827347
Jonasson Mattias Enar, Buecker Christa
What is an enhancer?
2023 BioEssays : news and reviews in molecular, cellular and developmental biology;45(10):e2300044.
PMID: 37256273
Thomas Henry Fabian, Buecker Christa
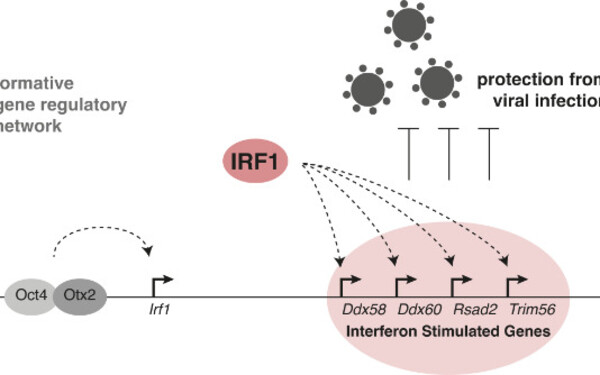
Transient upregulation of IRF1 during exit from naive pluripotency confers viral protection.
2022 EMBO reports;23(9):e55375.
PMID: 35852463
Romeike Merrit, Spach Stephanie, Huber Marie, Feng Songjie, Vainorius Gintautas, Elling Ulrich, Versteeg Gjis A, Buecker Christa
Temporal dissection of an enhancer cluster reveals distinct temporal and functional contributions of individual elements.
2021 Molecular cell;81(5):969, 982.e13, 969-982.e13.
PMID: 33482114
Thomas Henry F, Kotova Elena, Jayaram Swathi, Pilz Axel, Romeike Merrit, Lackner Andreas, Penz Thomas, Bock Christoph, Leeb Martin, Halbritter Florian, Wysocka Joanna, Buecker Christa
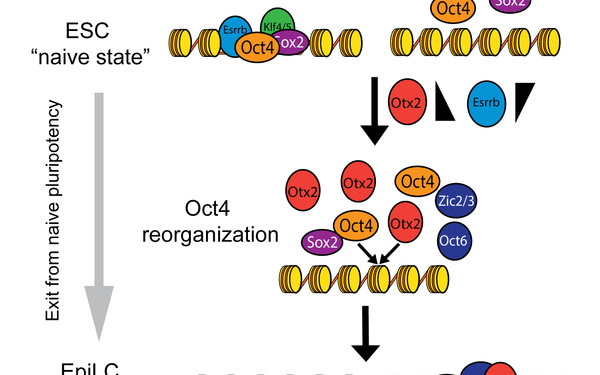
Reorganization of enhancer patterns in transition from naive to primed pluripotency.
2014 Cell stem cell(6)
PMID: 24905168
Buecker Christa, Srinivasan Rajini, Wu Zhixiang, Calo Eliezer, Acampora Dario, Faial Tiago, Simeone Antonio, Tan Minjia, Swigut Tomasz, Wysocka Joanna