MUT-7 exoribonuclease activity and localization are mediated by an ancient domain.
2024 Nucleic acids research;52(15):9076, 9091, 9076-9091.
PMID: 39188014
Busetto Virginia, Pshanichnaya Lizaveta, Lichtenberger Raffael, Hann Stephan, Ketting René F, Falk Sebastian
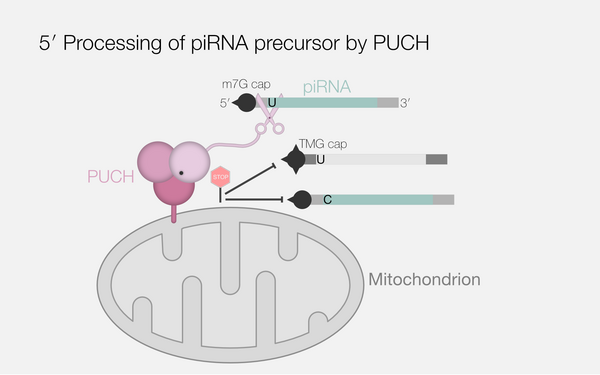
piRNA processing by a trimeric Schlafen-domain nuclease.
2023 Nature;622(7982):402, 409, 402-409.
PMID: 37758951
Podvalnaya Nadezda, Bronkhorst Alfred W, Lichtenberger Raffael, Hellmann Svenja, Nischwitz Emily, Falk Torben, Karaulanov Emil, Butter Falk, Falk Sebastian, Ketting René F
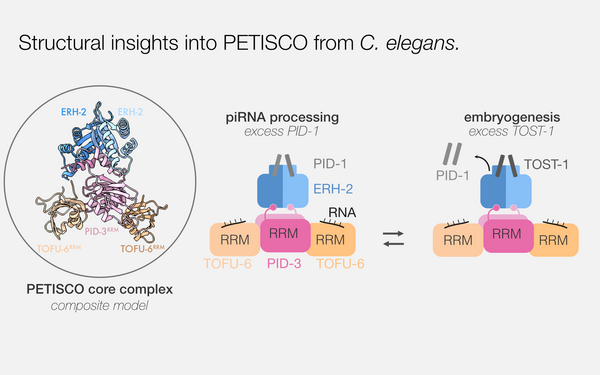
Structural basis of PETISCO complex assembly during piRNA biogenesis in .
2021 Genes & development;35(17-18):1304, 1323, 1304-1323.
PMID: 34413138
Perez-Borrajero Cecilia, Podvalnaya Nadezda, Holleis Kay, Lichtenberger Raffael, Karaulanov Emil, Simon Bernd, Basquin Jérôme, Hennig Janosch, Ketting René F, Falk Sebastian
A ribonuclease III involved in virulence of Mucorales fungi has evolved to cut exclusively single-stranded RNA.
2021 Nucleic acids research;49(9):5294, 5307, 5294-5307.
PMID: 33877360
Cánovas-Márquez José Tomás, Falk Sebastian, Nicolás Francisco E, Padmanabhan Subramanian, Zapata-Pérez Rubén, Sánchez-Ferrer Álvaro, Navarro Eusebio, Garre Victoriano
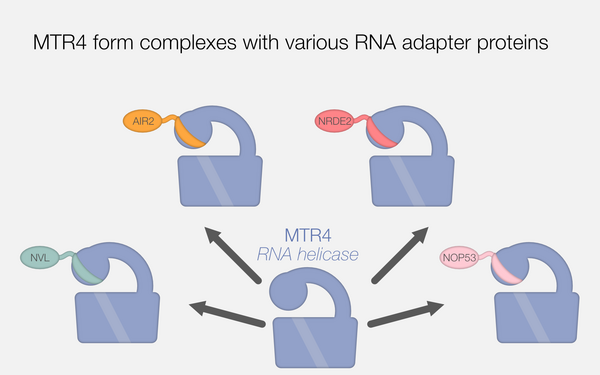
The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs.
2019 Nature communications;10(1):3393.
PMID: 31358741
Lingaraju Mahesh, Johnsen Dennis, Schlundt Andreas, Langer Lukas M, Basquin Jérôme, Sattler Michael, Heick Jensen Torben, Falk Sebastian, Conti Elena
Structure of the nuclear exosome captured on a maturing preribosome.
2018 Science (New York, N.Y.)(6385)
PMID: 29519915
Schuller Jan Michael, Falk Sebastian, Fromm Lisa, Hurt Ed, Conti Elena
Mpp6 Incorporation in the Nuclear Exosome Contributes to RNA Channeling through the Mtr4 Helicase.
2017 Cell reports(10)
PMID: 28877463
Falk Sebastian, Bonneau Fabien, Ebert Judith, Kögel Alexander, Conti Elena
Structural insights into the interaction of the nuclear exosome helicase Mtr4 with the preribosomal protein Nop53.
2017 RNA (New York, N.Y.)(12)
PMID: 28883156
Falk Sebastian, Tants Jan-Niklas, Basquin Jerôme, Thoms Matthias, Hurt Ed, Sattler Michael, Conti Elena
Structure of the RBM7-ZCCHC8 core of the NEXT complex reveals connections to splicing factors.
2016 Nature communications
PMID: 27905398
Falk Sebastian, Finogenova Ksenia, Melko Mireille, Benda Christian, Lykke-Andersen Søren, Jensen Torben Heick, Conti Elena
The molecular architecture of the TRAMP complex reveals the organization and interplay of its two catalytic activities.
2014 Molecular cell(6)
PMID: 25175027
Falk Sebastian, Weir John R, Hentschel Jendrik, Reichelt Peter, Bonneau Fabien, Conti Elena