The Question
Our mission is to identify and address fundamental problems in cell biology in this time and age when science and technology are rapidly advancing. We do this by looking at cells using high-resolution microscopy with a passion for new discoveries. In addition, we are committed to empowering the next generation of scientists by fostering independence, self-respect, and creativity, so that they can cultivate original ideas and drive groundbreaking inventions.
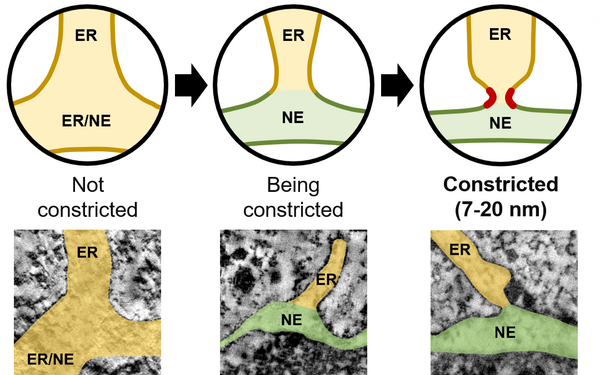
Eukaryotic cells contain many different organelles which enable them to compartmentalise biochemical and biological processes. The inter-organelle communication is tightly controlled to respond to environmental cues and coordinate homeostasis. A key organelle for the inter-organelle communication is the endoplasmic reticulum (ER), which is the site of lipid and membrane protein synthesis. The ER is directly connected to the nucleus by junctions with the outer nuclear membrane. This ER-to-nucleus connectivity is crucial for supplying the lipids and proteins that are synthesized in the ER to the nucleus. However, the biogenesis, architecture and maintenance of the ER-nucleus connection have remained elusive.