The Question
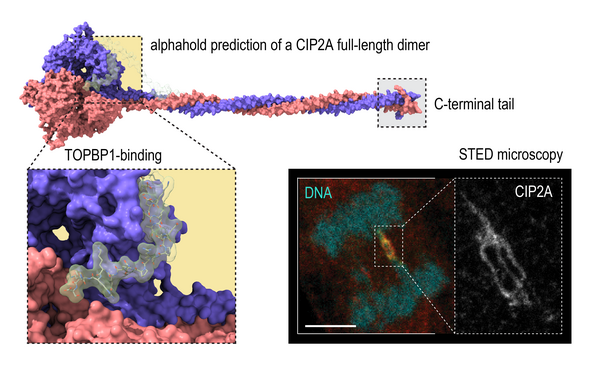
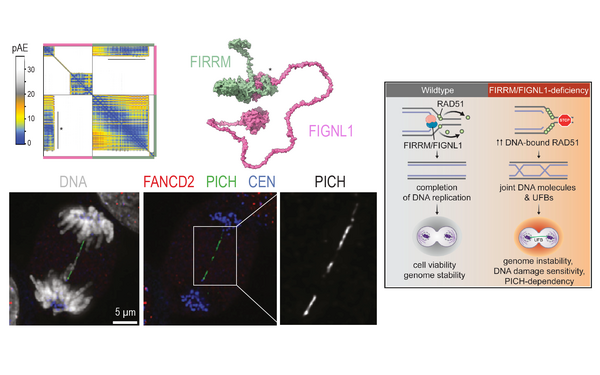
The propagation of life relies on the continuous replication and distribution of genomes. The Huis lab studies i) how DNA lesions are recognised and repaired during mitosis and ii) how DNA bridges are processed in anaphase. As sister chromatids segregate in anaphase, pieces of DNA that connect them are stretched into DNA bridges (see image). These bridges contain catenated, incompletely replicated, or damaged DNA and their resolution before cytokinesis is important to prevent the transmission of instable genomes to the arising daughter cells. We want to understand the protein machinery that safeguards our genomes during mitosis at a molecular and mechanistic level.
Image: an RPE1 cell in anaphase stained for DNA, PICH (yellow), and CIP2A (magenta). Courtesy of Panagiotis Martzios, PhD student in the Huis lab.